Quick start
IgPhyML is easiest to use when run indirectly through the R package Dowser. Most of these instructions require Dowser 1.0.0 or higher, and IgPhyML to be available in the Docker container or compiled locally.
The following commands should work as a first pass on many reasonably sized datasets, but if you really want to understand what’s going on or make sure what you’re doing makes sense, please check out the rest of the website.
Build trees and estimate model parameters
Open an R session (can type R into teriminal) in the Docker container:
#!/usr/bin/R
library(dowser)
library(airr)
library(tidyr)
library(dplyr)
# read in data after clonal clustering and germline reconstruction
data <- read_rearrangement("/usr/local/share/igphyml/examples/example.tsv")
# format clones for tree building (only 1 clone has >1 unique sequence)
clones <- formatClones(data)
# build tree using IgPhyML
# executable location will be different different if locally compiled
igphyml_exec <- "/usr/local/share/igphyml/src/igphyml"
trees <- getTrees(clones, build="igphyml", exec=igphyml_exec)
To visualize trees and print out parameter estimates, enter these commands:
# generate plot objects
plots <- plotTrees(trees)
# plot tree in console (might not work in Docker)
plots[[1]]
# plot trees as a pdf
treesToPDF(plots, file="igphyml_example_trees.pdf")
# save trees for later
saveRDS(trees, "trees.rds")
# look at parameters
print(gather(bind_rows(trees$parameters)))
# A tibble: 13 × 2
# key value
# <chr> <chr>
# 1 clone 1
# 2 nseq 2
# 3 nsite 106
# 4 tree_length 0.2865
# 5 lhood -147.7453
# 6 kappa_mle 1.9527
# 7 omega_mle 0.7074
# 8 wrc_2_mle 4.3296
# 9 gyw_0_mle 2.6478
#10 wa_1_mle 5.5308
#11 tw_0_mle 0.7451
#12 syc_2_mle -0.99
#13 grs_0_mle 0.2133
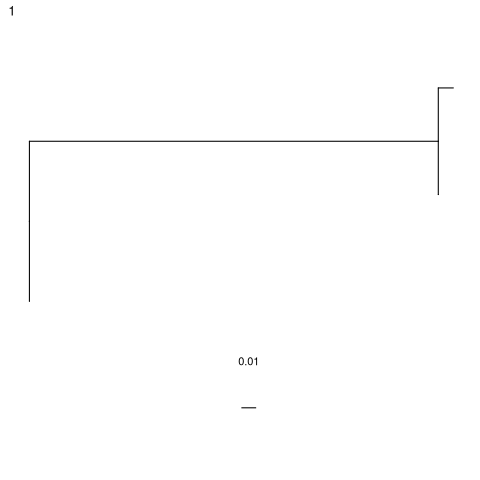
Lineage tree of example clone.
These commands process an AIRR-formatted dataset of BCR sequences that have been
clonally clustered
with germlines reconstructed.
It then quickly builds trees using the GY94 model and, using these
fixed topologies, estimates HLP19 model parameters. If analyzing multiple trees, this can be sped up by
setting the nproc option for formatClones and getTrees. Note that IgPhyML will run slowly when applied to large datasets,
so it may be worth down-sampling your data if you have clones larger than 100 unique sequences,
and removing clones with < 5 sequences if you have a many clones.
In addition to IgPhyML, Dowser offers many options for building trees and visualizing trees which may be more computationally efficient for larger datasets but don’t estimate the same potentially parameters as IgPhyML, such as omega (aka dN/dS or R/S).
To see more advanced options for IgPhyML tree building in Dowser, check out the documentation
for buildIghyml. You can pass any
option of this function to getTrees, e.g.
getTrees(clones, partition="cf", build="igphyml", exec=igphyml_exec) to estimate separate omegas for
CDRs and FWRs.